Research
How do environmental changes reshape microbial symbiosis?
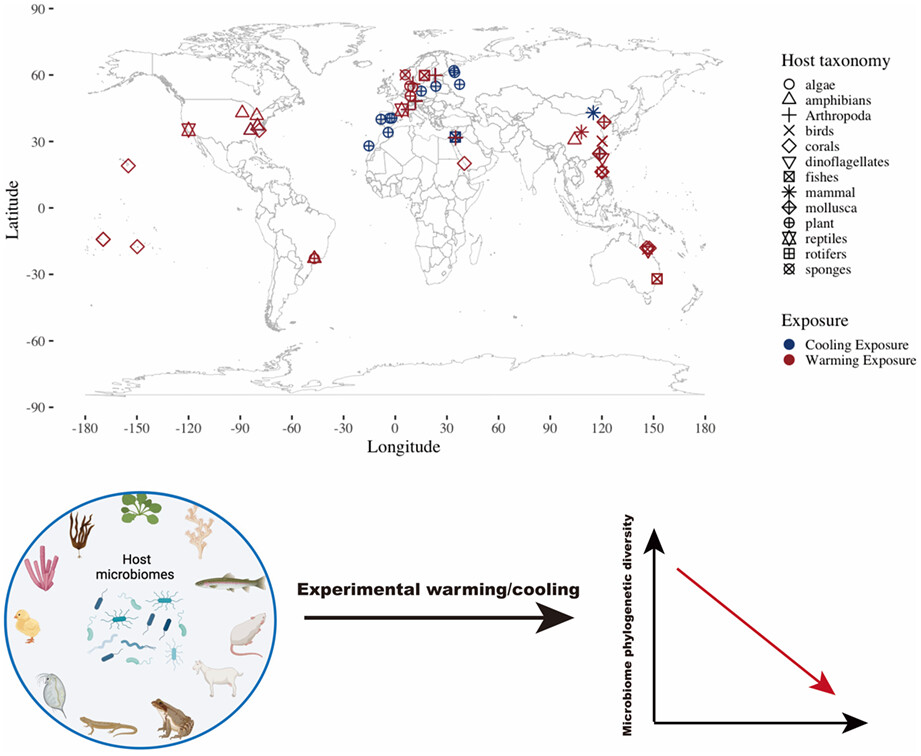
Microbial symbiosis is ubiquitous in nature, spanning the mutualism-to-parasitism continuum. Global climate change has led to more extreme thermal events. The extent to which non-pathogenic microbiome communities change in response to warming or cooling may be important for predicting host performance under global change (Li et al., 2025). Using a meta-analysis of 1377 microbiomes from 43 terrestrial and aquatic species, we found a decrease in the amplicon sequence variant-level microbiome phylogenetic diversity and alteration of microbiome composition under both experimental warming and cooling. We showed that the host habitat and experimental factors affected microbiome diversity and composition more than host biological traits (Li et al., 2023).
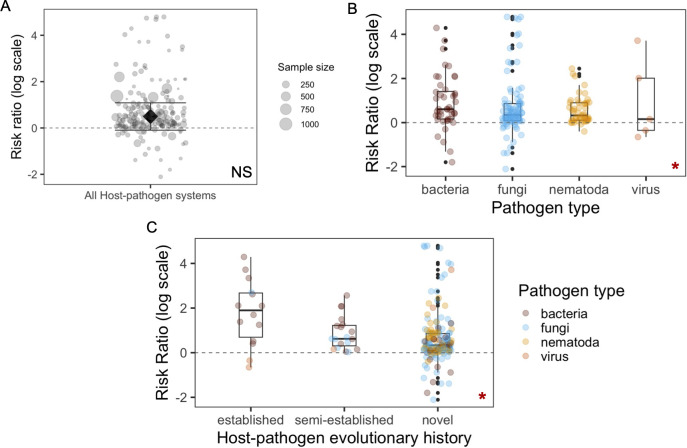
Warming events can sometimes lead to more infectious disease outbreaks, putting animal persistence at risk. The extent to which warming temperatures and infection may together impair host health is unclear. Using a meta-analysis of >190 effect sizes representing 101 ectothermic animal host–pathogen systems, we demonstrate that warming significantly increased the mortality of hosts infected by bacterial pathogens. Pathogens that have been evolutionarily established within the host species showed higher virulence under warmer temperatures. We found that compared to established infections, novel infections were more deadly at lower/baseline temperatures. We revealed that the virulence of fungal pathogens increased only when temperatures were shifted upwards towards the pathogen thermal optimum (Li et al., 2024).
Understanding how microbiome structures change in response to infection and warming may be important for forecasting host performance under global change. To move beyond pairwise host-microbiome or host-pathogen interactions, we evaluated alterations in the microbiomes of several wild Caenorhabditis elegans isolates spanning a range of latitudes, upon warming temperatures and infection by the parasite Leucobacter musarum. Using 16S rRNA sequencing, we found that microbiome diversity decreased, and dispersion increased over time, with the former being more prominent in uninfected adults and the latter aggravated by infection. Exposing infected hosts to warming did not have an additive destabilizing effect on their microbiomes. Moreover, warming during pre-adult development alleviated the destabilizing effects of infection on host microbiomes (Li et al., 2024).
How does warming impact animal resistance ecology and evolution?
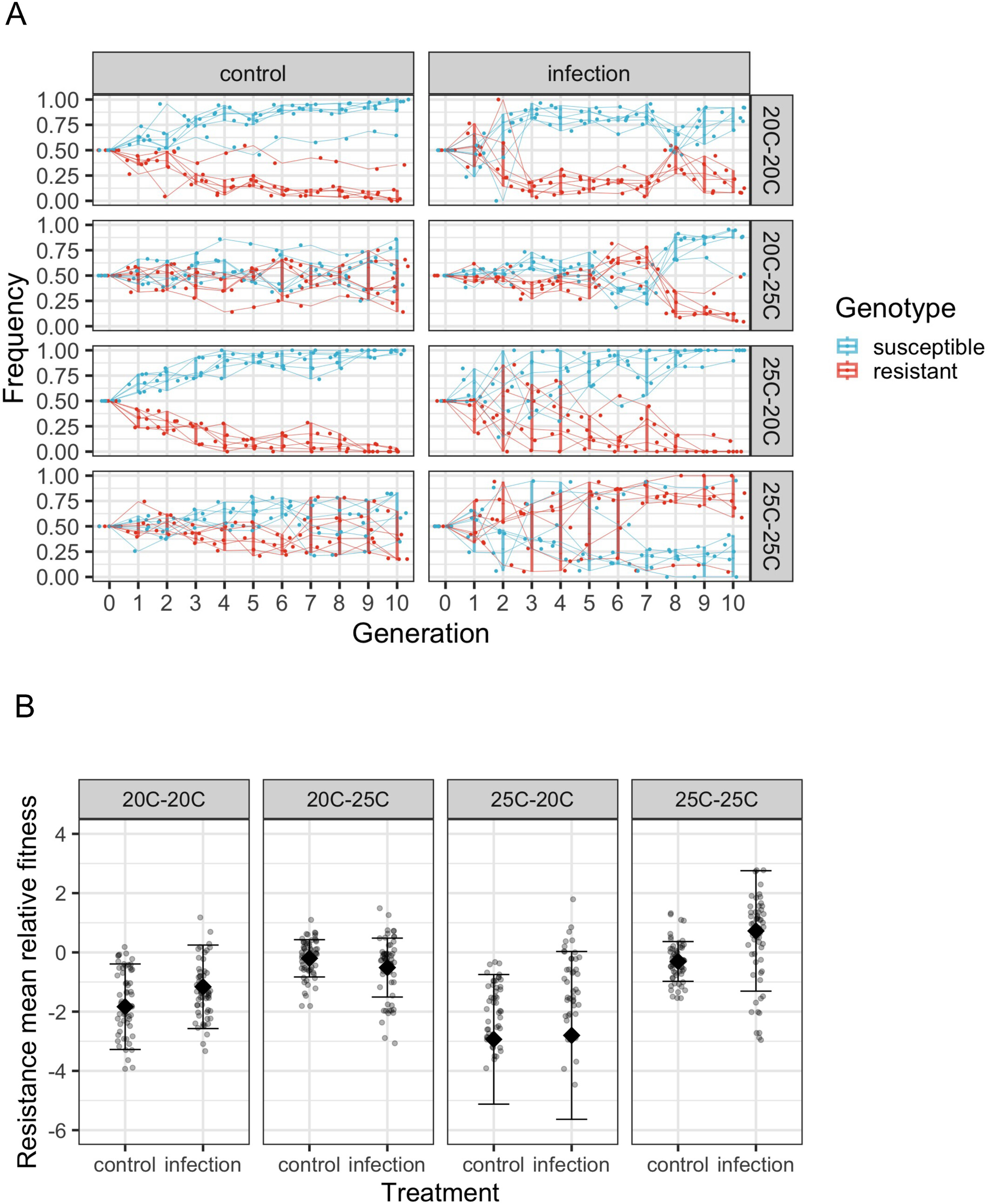
Climate change is increasing extreme heating events which may potentially exacerbate infectious diseases. Whether hosts can adapt to infection with rising temperatures is important for forecasting species persistence. We tested whether warming—at different host life stages—affects the ecological and evolutionary dynamics of resistance in Caenorhabditis elegans infected by a wild bacterial pathogen. We competed resistant and susceptible genotypes across 10 passages and tracked the spread of resistance in the population. Infection and prolonged warming strongly selected for the resistant genotype. Warming during host development induced plastic defences against infection, reducing the selective pressure for costly genetic-based resistance. Resistance was lost under ambient temperatures and periodic warming. Selection for resistance was likely weakened at ambient temperatures by the dilution effect, whereby the resistant genotype reduced pathogen transmission. Evolutionary dynamics of resistance depend on the balance among pathogen virulence, costs of genetic-based resistance, the dilution effect and plastic defences induced by temperature stress (Li et al., 2025).
What drives microbiome stability vs. flexibility in wild mammals and how these dynamics feed back onto host fitness?
While experimental systems are valuable for isolating microbial responses to specific perturbances, understanding the fitness consequences of microbiome dynamics in nature requires longitudinal studies at both individual and population levels in wild populations. Most existing work either captures population-level patterns at single timepoint, or relies on laboratory-maintained animals whose microbiomes may not reflect the ecological complexity and variability of natural environments. As a result, we still lack a clear understanding of the drivers of microbiome temporal dynamics, the conditions under which microbiome stability versus flexibility is beneficial, and how these dynamics translate into host fitness outcomes in the wild. Addressing this gap is critical for understanding how microbiome-mediated processes influence animal health and conservation outcomes in a rapidly changing world.
My postdoctoral research is currently focusing on addressing this gap by studying microbiome stability and variation in wild mammals, using polar bear as a model system. Polar bears are apex predators and key indicators of Arctic ecosystem health, with populations increasingly threatened by climate change. Polar bears typically experience extreme nutritional stress (prolonged fasting), especially during up to 8-month winter denning period for pregnant females. These conditions provide a natural experiment for identifying microbial taxa that persist under severe perturbation. These persistent microbial taxa may represent functionally or evolutionarily essential components of the host microbiome.
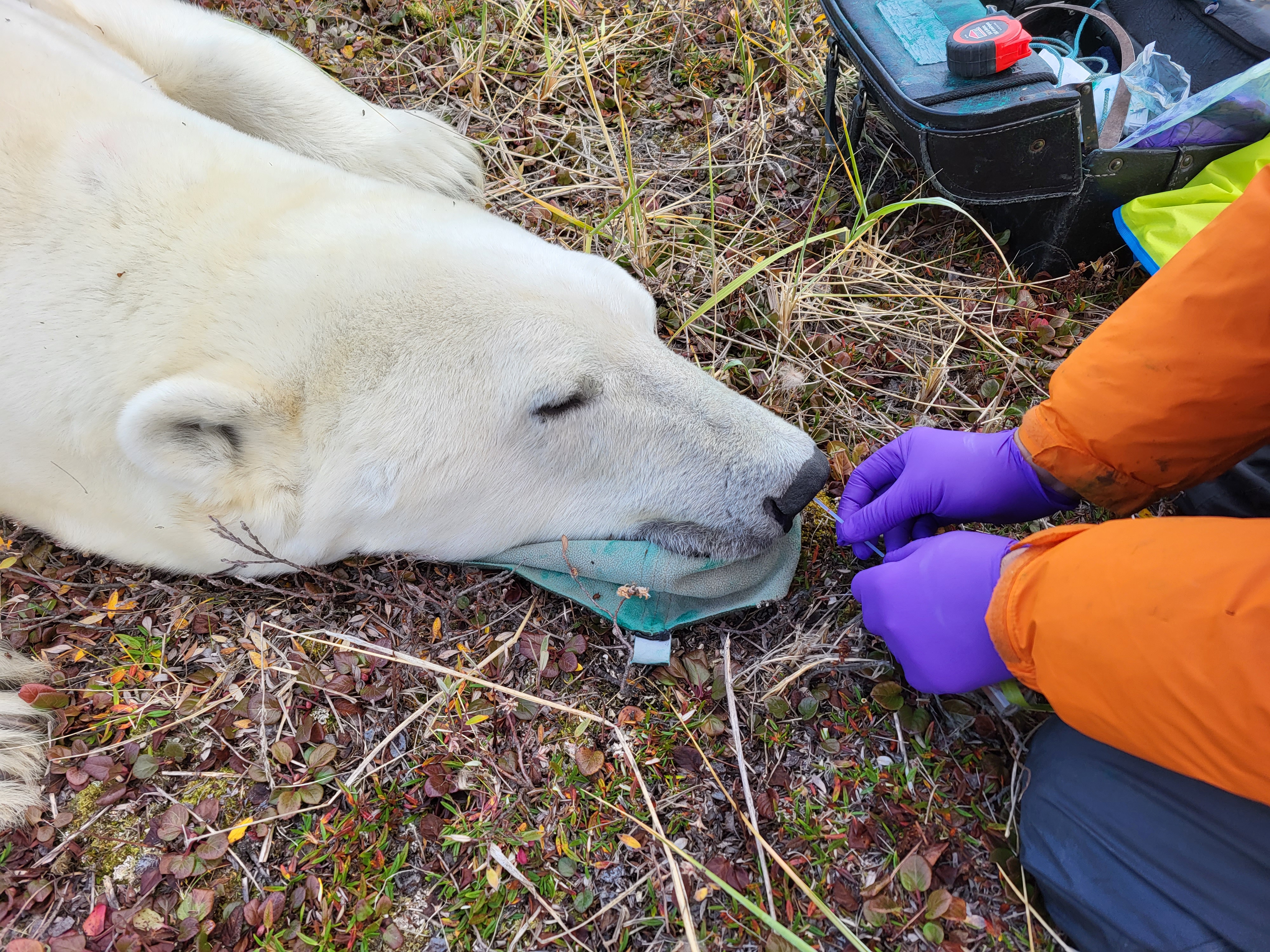
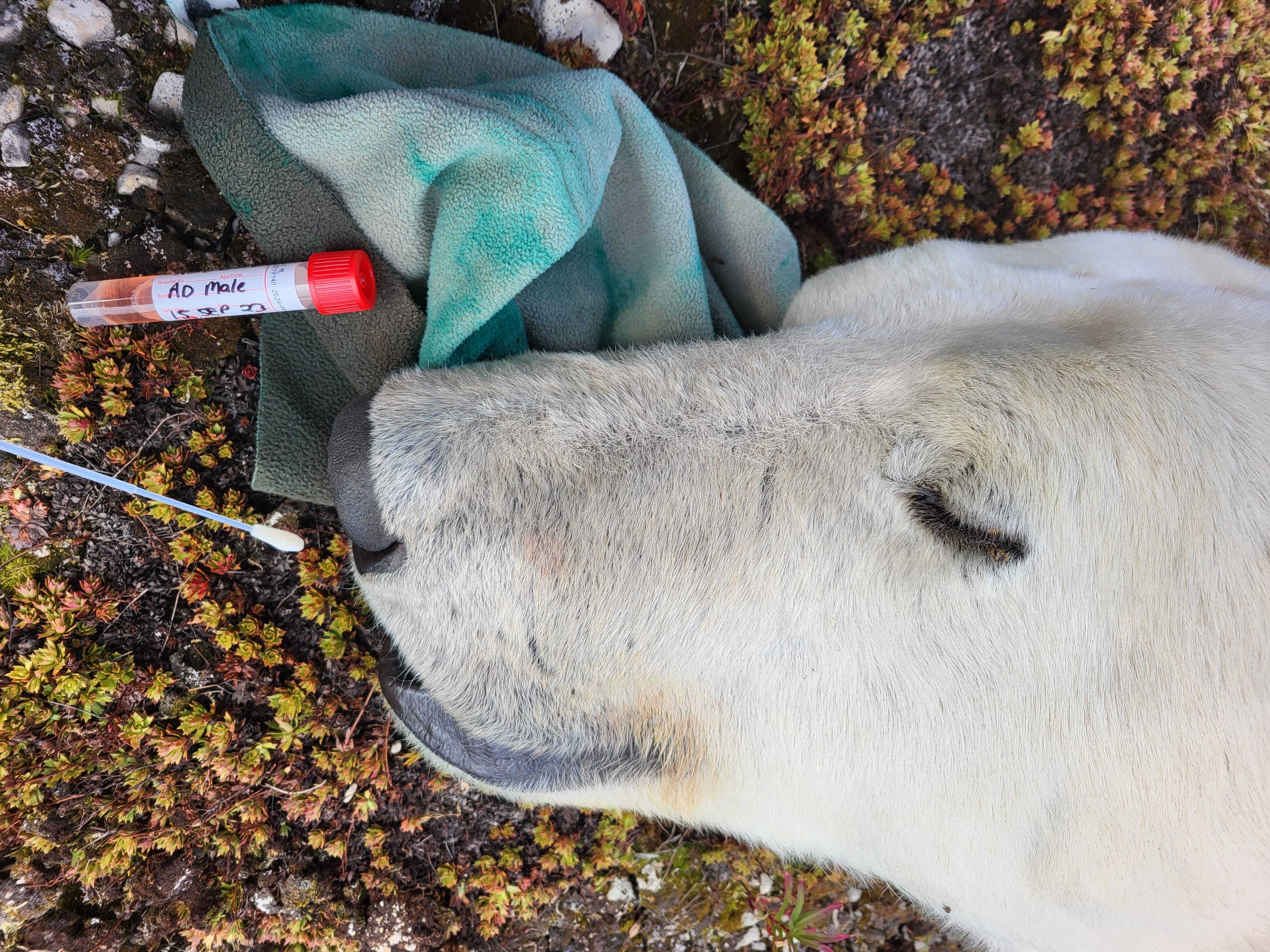
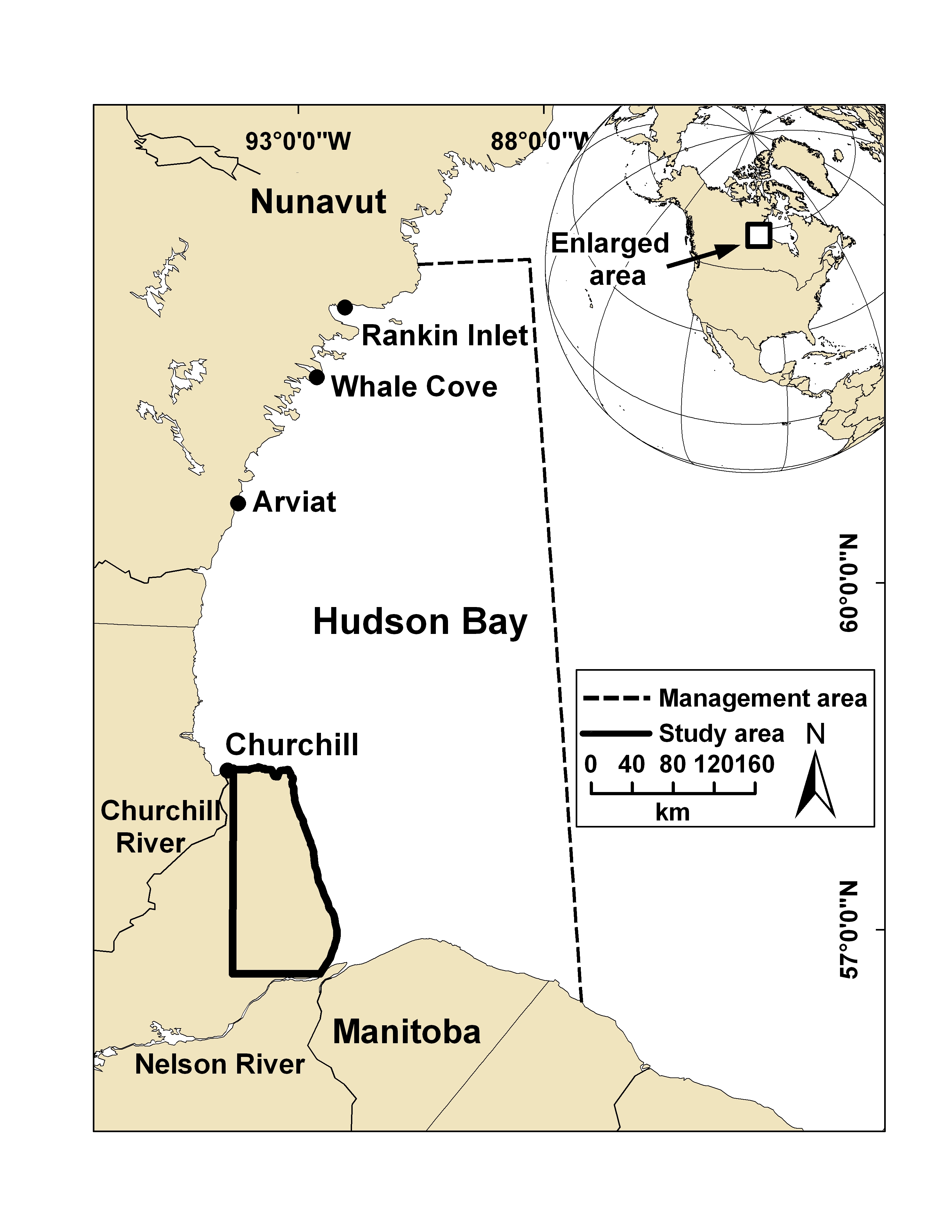


The Churchill polar bear population in Canada is among the most intensively monitored in the world. Through our collaboration, we have obtained microbiome samples (both rectal and nasal swabs) from over 200 bears across four timepoints over multiple years. Using metagenomics and metatranscriptomics, combined with detailed bear metadata, family and geographical information, I will first differentiate host-associated persistent vs. transient microbial taxa, and analyze their evolutionary histories and metabolic functions. We will further investigate how RNA viral infections disrupt microbial stability. Linking viral diversity to microbial restructuring and host body conditions will provide mechanistic insight into microbiome-mediated infection outcomes.
Stay tuned while this project is onging!
